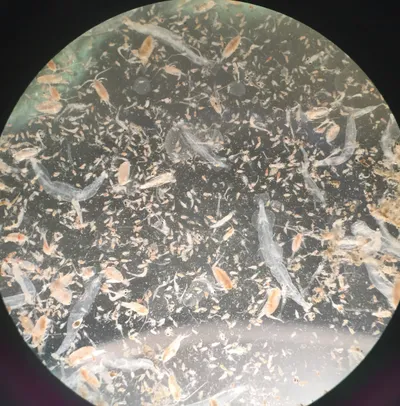
We're ocean detectives!
Have you ever wondered what lives deep in the ocean, where it's totally dark? We go there to find out! We study tiny animals that float in the sea and even tinier living things called microbes — they're so small you need a super-powered microscope to see them. We also use computers to read the special code inside every living thing (it's called DNA) to figure out who's who in the ocean. Think of us as underwater detectives, solving mysteries about creatures most people have never seen!
Ocean Tiny Creatures DNA Code Underwater Robots
Exploring hidden ocean life with DNA
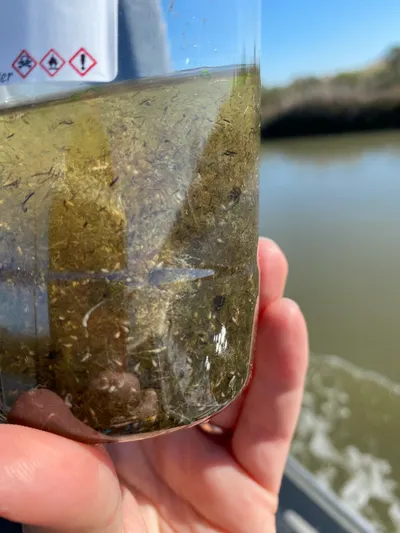
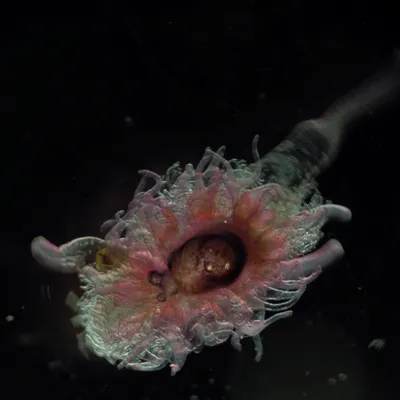
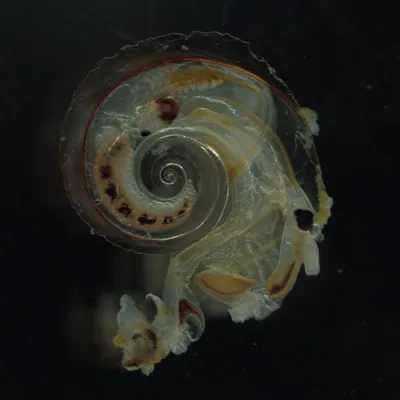
The Jungbluth Lab studies life in the ocean — from the zooplankton drifting near the surface to the microbes living deep beneath the seafloor. We use DNA to identify species that are hard to tell apart by sight, track what animals eat, and discover organisms that science hasn't described yet. Our fieldwork takes us to San Francisco Bay, the open Pacific, and even the deep sea using underwater robots. By combining biology with computer science, we build a clearer picture of how ocean ecosystems work and how they're changing.
Zooplankton Microbes DNA Barcoding Deep Sea Ecosystems
Marine ecology meets computational biology
The Jungbluth Lab bridges zooplankton ecology, microbial genomics, and computational biology. Michelle's group uses DNA barcoding and environmental DNA (eDNA) to study zooplankton communities, resolve food web interactions, and support conservation of threatened estuarine fishes like the longfin smelt. Sean's group applies metagenomics and bioinformatics to explore microbial and viral diversity in deep-sea and subsurface environments, while developing open-source tools and data standards that serve the broader research community. Together, we investigate life from the San Francisco Estuary to the deep ocean floor, combining fieldwork with large-scale data analysis.
eDNA Metagenomics Zooplankton Ecology Bioinformatics Food Webs
Integrating molecular ecology and environmental genomics
The Jungbluth Lab operates at the intersection of molecular ecology, environmental genomics, and computational biology. Michelle Jungbluth's research program applies qPCR-based quantification, DNA metabarcoding, and next-generation sequencing to characterize zooplankton population dynamics, resolve trophic interactions in estuarine food webs, and identify indicator species in human-impacted wetlands. Sean Jungbluth's work centers on metagenomic exploration of microbial and viral diversity in subsurface and deep-sea environments, with parallel efforts in AI/ML applications for environmental science and the development of community data standards (e.g., MIxS, MIEM). Current projects span the San Francisco Estuary, open-ocean pelagic systems, and deep biosphere sites, integrating field sampling with high-throughput sequencing and scalable bioinformatics pipelines.
Metagenomics qPCR Metabarcoding Subsurface Biosphere AI/ML Trophic Ecology